Linear regression (Stan)#
We will now conduct the same linear regression as in the previous chapter, but with Stan rather than PyMC. The Stan documentation has an example of linear regression model as well.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import arviz as az
from cmdstanpy import CmdStanModel
Data#
The data file can be downloaded here.
df = pd.read_csv('data/linreg.csv')
# Separating pre and post periods
df_train = df[df['period']=="pre"]
df_test = df[df['period']=="post"]
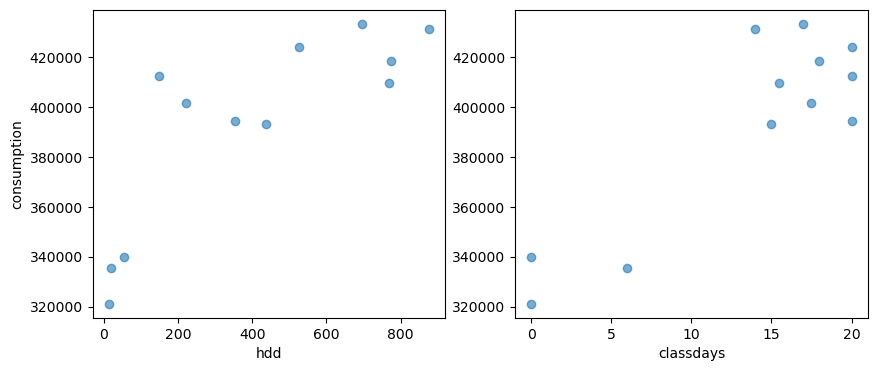
# A plot of the dependent variable vs. each of the explanatory variables
fig, axes = plt.subplots(1, 2, figsize=(10, 4))
axes[0].scatter(df_train['HDD'], df_train['consumption'], alpha=0.6)
axes[1].scatter(df_train['CLASSDAYS'], df_train['consumption'], alpha=0.6)
axes[0].set_ylabel("consumption")
axes[0].set_xlabel("hdd")
axes[1].set_xlabel("classdays")
Text(0.5, 0, 'classdays')
Model specification#
A Stan model is a block of text that can either be written in a separate file, or in the same script as the current code. Stan has high-level interfaces with most programming languages. Here, we’ll use CmdStanPy.
data {
int<lower=0> N; // number of data items
int<lower=0> K; // number of predictors
matrix[N, K] x; // predictor matrix
vector[N] y; // outcome vector
}
parameters {
real alpha; // intercept
vector[K] beta; // coefficients for predictors
real<lower=0> sigma; // error scale
}
model {
y ~ normal(x * beta + alpha, sigma); // likelihood
}
generated quantities {
array[N] real log_lik;
array[N] real y_hat;
for (j in 1:N) {
log_lik[j] = normal_lpdf(y[j] | x[j] * beta + alpha, sigma);
y_hat[j] = normal_rng(x[j] * beta + alpha, sigma);
}
}
This code should has been written into a dedicated file called linearregression.stan, which can be downloaded here.
model = CmdStanModel(stan_file='models/linearregression.stan')
If there is no compiled executable of this model in this filepath, this function will create one. If there is and the model has not been modified, it will load it without recompilation. This allows saving time when fitting the same model: compiling a new or modified model for the first time takes a while, but then the compiled executable is saved and the model may be re-used at will with different data sets.
Training#
The sample() method is used to do Bayesian inference over the model conditioned on data using using Hamiltonian Monte Carlo (HMC) sampling. It runs Stan’s HMC-NUTS sampler on the model and data and returns a CmdStanMCMC object. CmdStanPy allows specifying the data as a Python dictionary.
In the PyMC linear regression example, we fit two models: one with only HDD as explanatory variable, and one with also the class days. Let us do the same here:
# Data declaration for model 1
model1_data = {
"N": len(df_train),
"K": 1,
"x": df_train[['HDD']].values,
"y": df_train['consumption'].values,
}
# Fitting model 1
fit1 = model.sample(data=model1_data, show_progress=False)
09:54:13 - cmdstanpy - INFO - CmdStan start processing
09:54:13 - cmdstanpy - INFO - Chain [1] start processing
09:54:13 - cmdstanpy - INFO - Chain [2] start processing
09:54:13 - cmdstanpy - INFO - Chain [3] start processing
09:54:13 - cmdstanpy - INFO - Chain [4] start processing
09:54:14 - cmdstanpy - INFO - Chain [3] done processing
09:54:15 - cmdstanpy - INFO - Chain [1] done processing
09:54:15 - cmdstanpy - INFO - Chain [4] done processing
09:54:16 - cmdstanpy - INFO - Chain [2] done processing
# Data declaration for model 2
model2_data = {
"N": len(df_train),
"K": 2,
"x": df_train[['HDD', 'CLASSDAYS']].values,
"y": df_train['consumption'].values,
}
# Fitting model 2
fit2 = model.sample(data=model2_data, show_progress=False)
09:54:16 - cmdstanpy - INFO - CmdStan start processing
09:54:16 - cmdstanpy - INFO - Chain [1] start processing
09:54:16 - cmdstanpy - INFO - Chain [2] start processing
09:54:16 - cmdstanpy - INFO - Chain [3] start processing
09:54:16 - cmdstanpy - INFO - Chain [4] start processing
09:54:18 - cmdstanpy - INFO - Chain [3] done processing
09:54:19 - cmdstanpy - INFO - Chain [4] done processing
09:54:19 - cmdstanpy - INFO - Chain [1] done processing
09:54:19 - cmdstanpy - INFO - Chain [2] done processing
We can see than sampling is quite fast once the model has been compiled.
Note: the show_progress=False argument used above is only to simplify the display on this page. It can be removed or set to True to track the progress of sampling.
CmdStan has diagnostics utilities: stansummary and diagnose
The summary() method runs an utility that reads the outputs of all chains and computes summary statistics for all sampler and model parameters and quantities of interest. It also returns summaries of the total joint log-probability density lp__.
fs = fit1.summary(percentiles=(5, 50, 95))
fs.loc[['alpha', 'beta[1]', 'sigma', 'lp__']]
| Mean | MCSE | StdDev | 5% | 50% | 95% | N_Eff | N_Eff/s | R_hat | |
|---|---|---|---|---|---|---|---|---|---|
| alpha | 351957.0000 | 367.627000 | 13780.000000 | 329221.000 | 352384.0000 | 374190.000 | 1405.020000 | 730.259000 | 0.999853 |
| beta[1] | 100.2520 | 0.697350 | 26.573900 | 57.367 | 99.9952 | 143.995 | 1452.140000 | 754.750000 | 0.999405 |
| sigma | 27345.6132 | 203.997122 | 7403.755025 | 18267.100 | 25965.8000 | 41039.000 | 1317.212973 | 684.622127 | 1.000971 |
| lp__ | -117.5860 | 0.044984 | 1.419820 | -120.228 | -117.2350 | -116.031 | 996.235000 | 517.794000 | 1.002270 |
fs = fit2.summary(percentiles=(5, 50, 95))
fs.loc[['alpha', 'beta[1]', 'beta[2]', 'sigma', 'lp__']]
| Mean | MCSE | StdDev | 5% | 50% | 95% | N_Eff | N_Eff/s | R_hat | |
|---|---|---|---|---|---|---|---|---|---|
| alpha | 325605.00000 | 235.844000 | 9436.19000 | 310892.0000 | 325514.0000 | 340604.0000 | 1600.83000 | 1137.76000 | 1.004220 |
| beta[1] | 56.48890 | 0.383132 | 17.16070 | 29.7856 | 56.5464 | 83.7613 | 2006.19000 | 1425.86000 | 0.999786 |
| beta[2] | 3261.10000 | 17.712800 | 734.32600 | 2063.7100 | 3259.4900 | 4438.6500 | 1718.71000 | 1221.55000 | 1.002670 |
| sigma | 14078.61626 | 125.454950 | 4278.56431 | 9125.7000 | 13254.4000 | 21583.4000 | 1163.10931 | 826.65907 | 1.003890 |
| lp__ | -110.08700 | 0.062631 | 1.80114 | -113.5180 | -109.7060 | -108.0380 | 827.02500 | 587.79300 | 1.006430 |
Additionally, the diagnose() method runs an utility which analyzes the per-draw sampler parameters across all chains looking for potential problems which indicate that the sample isn’t a representative sample from the posterior.
print(fit2.diagnose())
Processing csv files: /tmp/tmplq8zu57i/linearregressionk24hs4cb/linearregression-20250923095416_1.csv, /tmp/tmplq8zu57i/linearregressionk24hs4cb/linearregression-20250923095416_2.csv, /tmp/tmplq8zu57i/linearregressionk24hs4cb/linearregression-20250923095416_3.csv, /tmp/tmplq8zu57i/linearregressionk24hs4cb/linearregression-20250923095416_4.csv
Checking sampler transitions treedepth.
Treedepth satisfactory for all transitions.
Checking sampler transitions for divergences.
No divergent transitions found.
Checking E-BFMI - sampler transitions HMC potential energy.
E-BFMI satisfactory.
Effective sample size satisfactory.
Split R-hat values satisfactory all parameters.
Processing complete, no problems detected.
Extracting results#
The fit1 and fit2 objects include methods to access the sampler outputs. They are listed on CmdStanPy’s documentation but we can mention two of them:
draws_pd(): returns a pandas.DataFrame over all columns in the Stan CSV file. The argument vars can be used to restrict this to one or more variables.draws_xr(): returns an xarray.Dataset which maps model variable names to their respective values.
res_pd = fit1.draws_pd()
res_xr = fit1.draws_xr()
res_pd.head()
| chain__ | iter__ | draw__ | lp__ | accept_stat__ | stepsize__ | treedepth__ | n_leapfrog__ | divergent__ | energy__ | ... | y_hat[3] | y_hat[4] | y_hat[5] | y_hat[6] | y_hat[7] | y_hat[8] | y_hat[9] | y_hat[10] | y_hat[11] | y_hat[12] | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1.0 | 1.0 | 1.0 | -118.238 | 0.971145 | 0.339103 | 3.0 | 7.0 | 0.0 | 118.663 | ... | 373191.0 | 408420.0 | 449295.0 | 444354.0 | 519855.0 | 458727.0 | 473612.0 | 422564.0 | 345776.0 | 357898.0 |
| 1 | 1.0 | 2.0 | 2.0 | -118.237 | 0.993430 | 0.339103 | 4.0 | 15.0 | 0.0 | 119.749 | ... | 355499.0 | 364834.0 | 377669.0 | 427364.0 | 470069.0 | 417393.0 | 413354.0 | 397723.0 | 398708.0 | 354786.0 |
| 2 | 1.0 | 3.0 | 3.0 | -117.646 | 0.985189 | 0.339103 | 3.0 | 7.0 | 0.0 | 118.872 | ... | 337474.0 | 403931.0 | 393499.0 | 427051.0 | 424879.0 | 438156.0 | 407038.0 | 403268.0 | 361106.0 | 359410.0 |
| 3 | 1.0 | 4.0 | 4.0 | -118.426 | 0.659121 | 0.339103 | 3.0 | 7.0 | 0.0 | 119.950 | ... | 364002.0 | 395530.0 | 387969.0 | 494433.0 | 485013.0 | 462230.0 | 418238.0 | 418811.0 | 447337.0 | 341333.0 |
| 4 | 1.0 | 5.0 | 5.0 | -122.657 | 0.941061 | 0.339103 | 3.0 | 15.0 | 0.0 | 123.148 | ... | 479027.0 | 311505.0 | 310124.0 | 368594.0 | 493771.0 | 488210.0 | 505653.0 | 423499.0 | 392101.0 | 304690.0 |
5 rows × 37 columns
The ArviZ package for exploratory analysis of Bayesian models can be used with CmdStanPy. To make the most out of ArviZ, it is recommended to convert the results to InferenceData. Doing so improves the res_xr xarray by assigning all variables to the right groups.
idata1 = az.from_cmdstanpy(
posterior = fit1,
posterior_predictive="y_hat",
observed_data={"y": df_train['consumption'].values},
constant_data={"x": df_train[['HDD']].values},
log_likelihood="log_lik",
dims={
"y_hat": ["observations"],
"log_lik": ["observations"],
"y": ["observations"],
"beta": ["features"]
},
)
idata2 = az.from_cmdstanpy(
posterior = fit2,
posterior_predictive="y_hat",
observed_data={"y": df_train['consumption'].values},
constant_data={"x": df_train[['HDD', 'CLASSDAYS']].values},
log_likelihood="log_lik",
dims={
"y_hat": ["observations"],
"log_lik": ["observations"],
"y": ["observations"],
"beta": ["features"]
},
)
idata2
-
<xarray.Dataset> Size: 136kB Dimensions: (chain: 4, draw: 1000, features: 2) Coordinates: * chain (chain) int64 32B 0 1 2 3 * draw (draw) int64 8kB 0 1 2 3 4 5 6 7 ... 993 994 995 996 997 998 999 * features (features) int64 16B 0 1 Data variables: alpha (chain, draw) float64 32kB 3.308e+05 3.232e+05 ... 3.237e+05 beta (chain, draw, features) float64 64kB 26.95 3.918e+03 ... 3.976e+03 sigma (chain, draw) float64 32kB 1.053e+04 1.226e+04 ... 1.716e+04 Attributes: created_at: 2025-09-23T07:54:26.529345+00:00 arviz_version: 0.22.0 inference_library: cmdstanpy inference_library_version: 1.2.5 -
<xarray.Dataset> Size: 392kB Dimensions: (chain: 4, draw: 1000, observations: 12) Coordinates: * chain (chain) int64 32B 0 1 2 3 * draw (draw) int64 8kB 0 1 2 3 4 5 6 ... 993 994 995 996 997 998 999 * observations (observations) int64 96B 0 1 2 3 4 5 6 7 8 9 10 11 Data variables: y_hat (chain, draw, observations) float64 384kB 3.307e+05 ... 3.3... Attributes: created_at: 2025-09-23T07:54:26.630012+00:00 arviz_version: 0.22.0 inference_library: cmdstanpy inference_library_version: 1.2.5 -
<xarray.Dataset> Size: 392kB Dimensions: (chain: 4, draw: 1000, observations: 12) Coordinates: * chain (chain) int64 32B 0 1 2 3 * draw (draw) int64 8kB 0 1 2 3 4 5 6 ... 993 994 995 996 997 998 999 * observations (observations) int64 96B 0 1 2 3 4 5 6 7 8 9 10 11 Data variables: log_lik (chain, draw, observations) float64 384kB -10.64 ... -10.99 Attributes: created_at: 2025-09-23T07:54:26.666022+00:00 arviz_version: 0.22.0 inference_library: cmdstanpy inference_library_version: 1.2.5 -
<xarray.Dataset> Size: 204kB Dimensions: (chain: 4, draw: 1000) Coordinates: * chain (chain) int64 32B 0 1 2 3 * draw (draw) int64 8kB 0 1 2 3 4 5 6 ... 994 995 996 997 998 999 Data variables: lp (chain, draw) float64 32kB -110.9 -108.5 ... -108.6 -109.9 acceptance_rate (chain, draw) float64 32kB 1.0 0.9958 ... 0.9849 0.9353 step_size (chain, draw) float64 32kB 0.1959 0.1959 ... 0.2104 0.2104 tree_depth (chain, draw) int64 32kB 2 3 3 4 4 3 3 4 ... 4 4 5 3 3 4 4 n_steps (chain, draw) int64 32kB 3 7 11 31 15 11 ... 31 15 15 15 15 diverging (chain, draw) bool 4kB False False False ... False False energy (chain, draw) float64 32kB 116.9 111.3 ... 109.9 112.2 Attributes: created_at: 2025-09-23T07:54:26.586935+00:00 arviz_version: 0.22.0 inference_library: cmdstanpy inference_library_version: 1.2.5 -
<xarray.Dataset> Size: 192B Dimensions: (observations: 12) Coordinates: * observations (observations) int64 96B 0 1 2 3 4 5 6 7 8 9 10 11 Data variables: y (observations) int64 96B 321120 335520 ... 401760 339840 Attributes: created_at: 2025-09-23T07:54:26.640046+00:00 arviz_version: 0.22.0 inference_library: cmdstanpy inference_library_version: 1.2.5 -
<xarray.Dataset> Size: 304B Dimensions: (x_dim_0: 12, x_dim_1: 2) Coordinates: * x_dim_0 (x_dim_0) int64 96B 0 1 2 3 4 5 6 7 8 9 10 11 * x_dim_1 (x_dim_1) int64 16B 0 1 Data variables: x (x_dim_0, x_dim_1) float64 192B 12.7 0.0 19.4 6.0 ... 17.5 54.2 0.0 Attributes: created_at: 2025-09-23T07:54:26.652370+00:00 arviz_version: 0.22.0 inference_library: cmdstanpy inference_library_version: 1.2.5
We can now use the same posterior plots as we did with the PyMC output:
az.plot_trace(idata1)
array([[<Axes: title={'center': 'alpha'}>,
<Axes: title={'center': 'alpha'}>],
[<Axes: title={'center': 'beta'}>,
<Axes: title={'center': 'beta'}>],
[<Axes: title={'center': 'sigma'}>,
<Axes: title={'center': 'sigma'}>]], dtype=object)
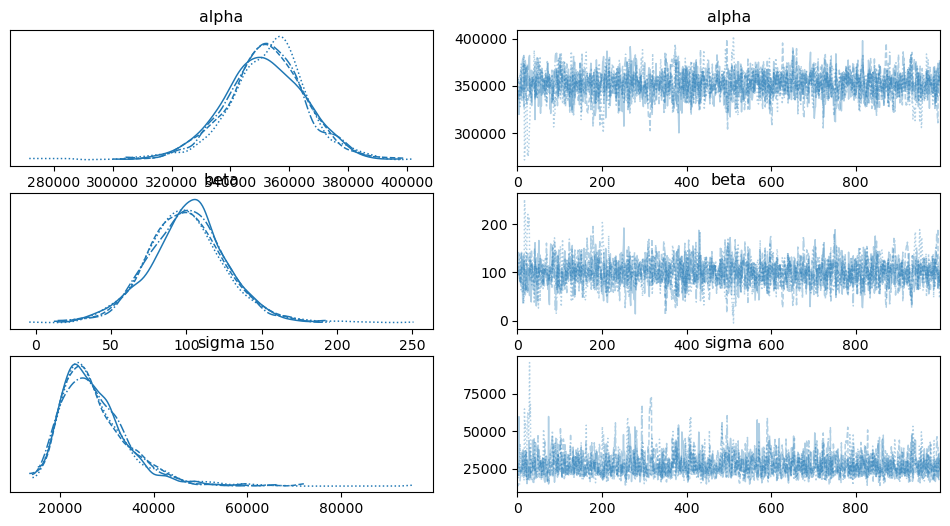
And here is the posterior predictive distribution:
y_post = idata2.posterior_predictive["y_hat"]
fig, axes = plt.subplots(1, 2, figsize=(10, 4))
axes[0].scatter(df_train['HDD'], df_train['consumption'], alpha=0.6, label='data')
axes[0].scatter(df_train['HDD'], y_post.mean(("chain", "draw")), c="C1", alpha=0.6, label='posterior mean')
az.plot_hdi(df_train['HDD'], y_post, ax=axes[0])
axes[0].set_xlabel("hdd")
axes[0].set_ylabel("consumption")
axes[0].legend()
axes[1].scatter(df_train['CLASSDAYS'], df_train['consumption'], alpha=0.6)
axes[1].scatter(df_train['CLASSDAYS'], y_post.mean(("chain", "draw")), c="C1", alpha=0.6)
az.plot_hdi(df_train['CLASSDAYS'], y_post, ax=axes[1])
axes[1].set_xlabel("class days")
Text(0.5, 0, 'class days')
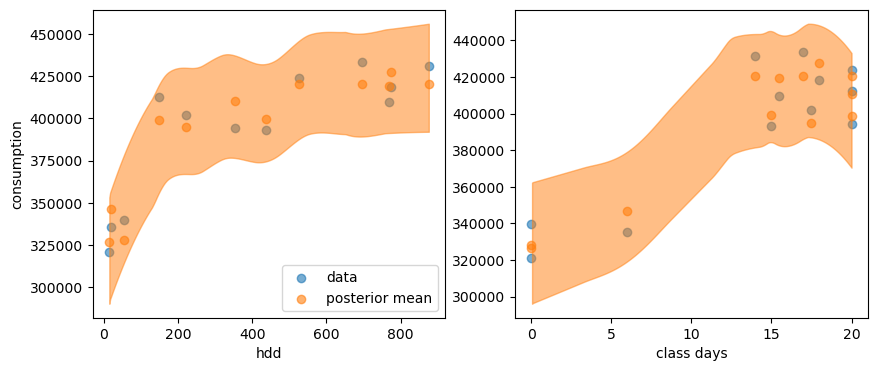
Reassuringly, this looks exactly like the posterior distribution we had from PyMC.
Model comparison metrics#
The built-in LOO method of ArviZ can be used now (see the previous chapter for a reminder of what is happening here):
df_comp_loo = az.compare({"model1": idata1, "model2": idata2}, ic="loo")
df_comp_loo
/home/simon/anaconda3/envs/bayes_env/lib/python3.11/site-packages/arviz/stats/stats.py:797: UserWarning: Estimated shape parameter of Pareto distribution is greater than 0.70 for one or more samples. You should consider using a more robust model, this is because importance sampling is less likely to work well if the marginal posterior and LOO posterior are very different. This is more likely to happen with a non-robust model and highly influential observations.
warnings.warn(
| rank | elpd_loo | p_loo | elpd_diff | weight | se | dse | warning | scale | |
|---|---|---|---|---|---|---|---|---|---|
| model2 | 0 | -132.533091 | 3.046308 | 0.000000 | 1.0 | 1.155242 | 0.000000 | True | log |
| model1 | 1 | -140.311554 | 2.490824 | 7.778464 | 0.0 | 2.074431 | 2.016874 | False | log |
